A26 - Modelling mechanisms of membrane shaping, fusion, and fission in endo- and exocytosis by protein scaffolds
Principal Investigator
Prof. Dr. Michael Kozlov, Tel Aviv University, Israel
Project A26 addresses, theoretically, different physical mechanisms of mechanical processes and transformations occurring with and within living cells. Over that last three decades, our group has been involved in reaching some understanding of the physical mechanisms underlying membrane shaping and remodeling by proteins.
Scaffolding mechanism
For instance, we have investigated how membrane curvature is created by different hydrophilic proteins, which bind the surface of one membrane monolayer leading to membrane asymmetry and, hence, development of membrane curvature. The results of our computational analysis of membrane shaping by an arc-like membrane scaffold imitating N-BAR dimer including the membrane-mediated interaction between the scaffolds are presented in (Schweitzer & Kozlov,2015; Figure 1).
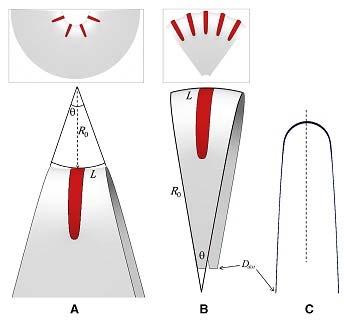
Figure 1: Example of computational results on membrane shaping by crescent-like protein scaffolds. (A) Sheet with negative curvature of the edge line (top) and its elementary section (bottom). (B) Sheet with positive curvature of the edge line, (top) and its elementary section (bottom). (C) Side view of a typical sheet section. Each section has mirror updown symmetry; (dashed line) symmetry plane. (Schweitzer & Kozlov, 2015)
Hydrophobic insertion mechanism
The hydrophobic insertion (or wedging) mechanism, is based on shallow embedding in one of the membrane leaflets of a hydrophobic or amphipathic protein domain (Zimmerberg & Kozlov, 2006), such as N-terminal amphipathic helices (Ford et al., 2002) or short hydrophobic loops (McMahon et al., 2010). Shallow protein insertions have been demonstrated (Ford et al., 2002) and predicted (Campelo et al., 2008; Fuhrmans & Marrink, 2012) to be powerful generators of membrane curvature. An amphipathic protein domain penetrating the external lipid monolayer only up to the interface between the lipid polar heads and the hydrocarbon tails pushes aside the polar heads while leaving the area underneath the insertion to be filled by splayed and tilted acyl chains. This results in a strong local asymmetry in the monolayer structure, which generates considerable local membrane curvatures. The shallower insertions are embedded per unit area of the membrane surface, the greater will be the global membrane curvature in the region occupied by the insertions (Campelo et al., 2008; Figure 2).
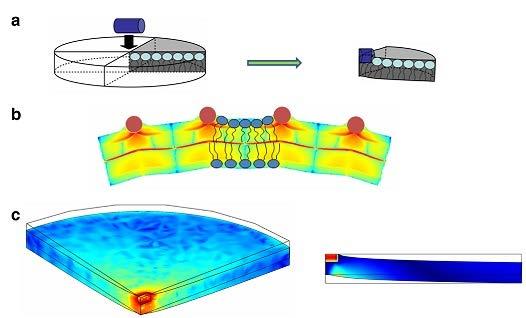
Figure 2: Hydrophobic insertion mechanism of membrane bending. Results of computational modelling of membrane curvature generation by rod-like particles imitating amphipathic helices 1, 2. The computations
were performed by COMSOL Multiphysics software. (a) Insertion of a helix. (b) Computational result for membrane bending by rows of rods. The figure shows an element of cross-section of a resulting tubule-like membrane. (c) Bending of a membrane element by a single insertion. Left panel – a 3D view, right panel – a cross-section element.
Modelling membrane fission
Membrane fusion and fission involve, at the intermediate stages of the reaction, drastic although local changes in membrane structure (Figure 3). Our group worked over decades on the “stalk” mechanism of membrane fusion and fission which by now is an established mechanism of the initial stages of membrane remodelling (see for review Chernomordik & Kozlov, 2005). Based on the stalk hypothesis, we developed a model for the entire pathway of membrane fission beginning from the constricted membrane neck (Figure 3A) and proceeding via a hemifission intermediate (Figure 3B) to complete membrane separation. The membrane stalk in this case isa lipidic connection formed as a result of fission of the internal monolayer of the neck preceding fission of the external monolayer (Figure 3B insert). We determined the structure and energy of the stalk intermediate by optimizing the packing of the hydrocarbon and the shape of the lipid monolayer surfaces (Kozlovsky & Kozlov, 2002).
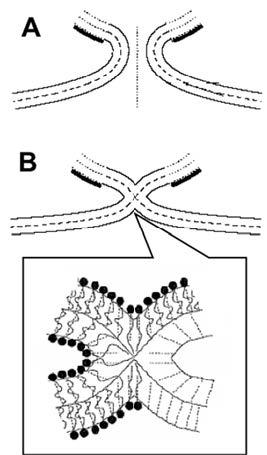
Figure 3: Computational results for pathway of membrane fission. (A) Membrane neck under mechanical stress. (B) Fission stalk. (Kozlovsky & Kozlov, 2003)
Currently, we also investigate the mechanisms of action of several specific proteins and protein complexes in clathrin‐mediated endocytosis (CME), clathrin independent endocytosis (CIE), and synaptic vesicle fusion with plasma membrane in the active zone (AZ). The goal of the modeling is to provide testable predictions on the protein structure transformations and protein‐membrane interactions, which drive membrane shaping and remodeling in the course of endocytic vesicle formation, in collaboration with the experimental projects of Haucke/ Maritzen (A01), Freund/ Haucke (A07) and Daumke (A11), and the computational project of Noe (A04). Together with the experimental project of Sigrist (A03) we study synaptic vesicle release and fusion with the plasma membrane of the AZ. Specifically, we analyze, computationally, the scenario of spatio‐temporal interplay between several proteins known to be involved in CME.
References:
- Haimov E, Urbakh M, Kozlov MM. Negative tension controls stability and structure of intermediate filament networks. Sci Rep. 12(1):16 (2022)
- Zucker B, Kozlov MM. Mechanism of shaping membrane nanostructures of endoplasmic reticulum. Proc Natl Acad Sci U S A. 119(1):e2116142119 (2022)
- Wang N, Clark LD, Gao Y, Kozlov MM, Shemesh T, Rapoport TA. Mechanism of membrane-curvature generation by ER-tubule shaping proteins. Nat Commun. 12(1):568 (2021)
- Golani G, Leikina E, Melikov K, Whitlock JM, Gamage DG, Luoma-Overstreet G, Millay DP, Kozlov MM, Chernomordik LV. Myomerger promotes fusion pore by elastic coupling between proximal membrane leaflets and hemifusion diaphragm. Nat Commun. 12(1):495 (2021)
- Ben Zucker, Gonen Golani, Michael M. Kozlov. Model for ring closure in ER tubular network dynamics. bioRxiv 2021.11.18.469198
- Haimov E, Windoffer R, Leube RE, Urbakh M, Kozlov MM. Model for Bundling of Keratin Intermediate Filaments. Biophys J. 119(1):65-74 (2020)
- Parton RG, Kozlov MM, Ariotti N. Caveolae and lipid sorting: Shaping the cellular response to stress. J Cell Biol. 219(4):e201905071 (2020)
- Choudhary, V., G. Golani, A.S. Joshi, S. Cottier, R. Schneiter, W.A. Prinz, and M.M. Kozlov. Architecture of Lipid Droplets in Endoplasmic Reticulum Is Determined by Phospholipid Intrinsic Curvature. Curr Biol. 28(6),915-926 (2018)
- Wu, X.S., S. Elias, H. Liu, J. Heureaux, P.J. Wen, A.P. Liu, M.M. Kozlov, and L.G. Wu. Membrane Tension Inhibits Rapid and Slow Endocytosis in Secretory Cells. Biophys. J. 113 (11), 2406-2414 (2017)
- Schweitzer, Y., and M.M. Kozlov. Membrane-Mediated Interaction between Strongly Anisotropic Protein Scaffolds. PLoS Comp. Biol. 11(2): 2 e1004054 (2015)
- Schweitzer, Y., T. Shemesh, and M.M. Kozlov. A Model for Shaping Membrane Sheets by Protein Scaffolds. Biophys. J. 109(3): 564-573 (2015)
- Terasaki, M., T. Shemesh, N. Kasthuri, R.W. Klemm, R. Schalek, K.J. Hayworth, A.R. Hand, M. Yankova, G. Huber, J.W. Lichtman, T.A. Rapoport, and M.M. Kozlov. Stacked endoplasmic reticulum sheets are connected by helicoidal membrane motifs. Cell. 154:285-96 (2013)
- Elia, N., G. Fabrikant, M.M. Kozlov, J. Lippincott-Schwartz. Computational model for cytokinetic abscission driven by ESCRT-III polymerization and remodeling. Biophys. J. 102: 2309-2320 (2012)
- Boucrot, E., A. Pick, G. Camdere, N. Liska, E. Evergren, H.T. McMahon, and M.M. Kozlov. Hydrophobic insertions promote, while crescent BAR scaffolds limit vesicle membrane fission. Cell. 149: 124-136 (2012)
- Shibata, Y., T. Shemesh, W.A. Prinz, A.F. Palazzo, M.M. Kozlov, and T.A. Rapoport. Mechanisms determining the morphology of the peripheral ER. Cell. 143:774-88 (2010)
- McMahon, H.T., M.M. Kozlov, and S. Martens. Membrane curvature in synaptic vesicle fusion and beyond. Cell. 140:601-605 (2010)
- Campelo, F., G. Fabrikant, H.T. McMahon, and M.M. Kozlov. Modeling membrane shaping by proteins: focus on EHD2 and N-BAR domains. FEBS Lett. 584:1830-9 (2010)