Z02 - Advanced and super-resolution imaging to resolve the nanoscale structure and dynamics of membrane scaffolds
Principal Investigator
Prof. Dr. Helge Ewers, FU Berlin
Project Z02 (Schmoranzer/Ewers) provides both the infrastructure as well as the expertise to apply advanced imaging technology towards the resolution of spatiotemporal organization of membrane scaffolds that are hidden behind the diffraction barrier. In the past, the most commonly used super‐resolution methods, including SIM (structured illumination), STED (stimulated emission depletion) and dSTORM (direct stochastic optical reconstruction), have been established and successfully applied toward the aims of the A projects.
Further improvement of these methods (Lehmann et al., 2015; Lehmann et al., 2016) enabled us to resolve nanoscale details of membrane scaffolds within cell cultures, neurons and tissues (Albrecht et al., 2016; Gimber et al., 2015; Koo et al., 2015; Wetzel et al., 2016; Schöneberg et al., 2017).
Furthermore, we developed a number of novel imaging techniques such as a combination of expansion microscopy with STED (Gao et al., 2018) with nanobody‐based labeling of microtubules (Mikhaylova et al., 2015) and the multicolor single-molecule localization approach SD-DNA-PAINT (spectral demixing DNA point accumulation for imaging in nanoscale topography, Gimber et al. 2022).
Currently, the Schmoranzer lab is focusing on the application of their novel super-resolution methods to several biological projects within the SFB958 and on the implementation of more than three colors to SD-DNA-PAINT.
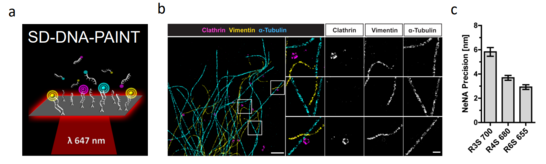
Figure 1: Spectral Demixing (SD) DNA-PAINT. (a) Principle of SD-DNA-PAINT. The transient binding of freely diffusing fluorescently labeled oligonucleotides to complementary DNA oligonucleotides that are linked to the target structure produces the required blinking for single molecule localization. Multiple dyes are excited by a single laser line and the mixed emission is split via a dichroic mirror on two sides of a camera. Localizations detected in both channels are paired and demixed via the channel-specific intensity values. (b) Example of a rendered triple-color SD-DNA-PAINT image of clathrin coated vesicles (magenta), vimentin (yellow) and microtubules (cyan). (d) Triple color SD-DNA-PAINT has a localization precision of 3-6 nm.
The Ewers lab focuses on (i) further developing alternative methods to enhance resolution (ExSTED) and (iI) establishing quantitative single molecule localization methods (DNA‐qPAINT).
We have further developed the Expansion microscopy through combining it with superresolution stimulated emission depletion (STED) (Figure 2). This new application (ExSTED) yields a very high resolution of up to 8 nm in xy and up to 45 nm isotropically in xyz. To further increase resolution we plan to develop novel labeling schemes and to combine ExSTED with novel Expansion microscopy approaches that allow an expansion of gels of up to 10- to 20-fold. This technique is now being implemented in several projects of the consortium (i.e. A22).
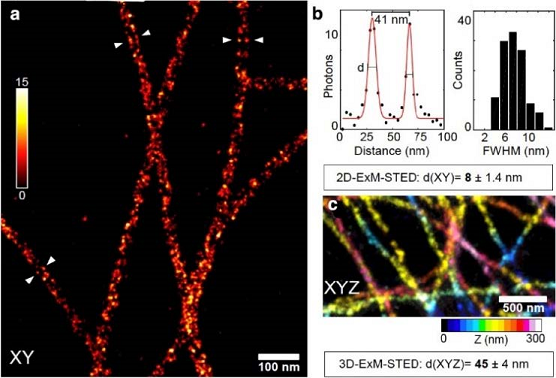
Figure 2: Expansion stimulated emission depletion microscopy (ExSTED). (a) ExSTED fluorescence micrograph of several microtubules (MTs). The MTs exhibit a « railroad-track » pattern of pairwise aligned filaments, demonstrating the hollow nature of the MTs which can be resolved using this method. (b) A quantification of the distance between the lines to the left and right of the MTs results in a full-width at half maximum of < 10 nm. (c) Part of a 3D projection of cellular MTs color-coded for Z-dimension. In 3D ExSTED an isotropic resolution of below 50 nm is possible.
References:
- Schmerl B, Gimber N, Kuropka B, Stumpf A, Rentsch J, Kunde SA, von Sivers J, Ewers H, Schmitz D, Freund C, Schmoranzer J, Rademacher N, Shoichet SA. The synaptic scaffold protein MPP2 interacts with GABAA receptors at the periphery of the postsynaptic density of glutamatergic synapses. PLoS Biol. 20(3) (2022)
- Gimber, N. Strauss, S. Jungmann, R. Schmoranzer, J. Simultaneous multicolor DNA-PAINT without sequential fluid exchange using spectral demixing. Nano Letters (2022) In Press.
- Voss M, Kleinau G, Gimber N, Janek K, Bredow C, Thery F, Impens F, Schmoranzer J, Scheerer P, Kloetzel PM, Beling A. A cytosolic disulfide bridge-supported dimerization is crucial for stability and cellular distribution of Coxsackievirus B3 protein 3A. FEBS J. (2022)
- Wilhelmi I, Grunwald S, Gimber N, Popp O, Dittmar G, Arumughan A, Wanker EE, Laeger T, Schmoranzer J, Daumke O, Schürmann A. The ARFRP1-dependent Golgi scaffolding protein GOPC is required for insulin secretion from pancreatic β-cells. Mol Metab. 45:101151 (2021)
- Lee CY, Petkova M, Morales-Gonzalez S, Gimber N, Schmoranzer J, Meisel A, Böhmerle W, Stenzel W, Schuelke M, Schwarz JM. A spontaneous missense mutation in the chromodomain helicase DNA-binding protein 8 (CHD8) gene: a novel association with congenital myasthenic syndrome. Neuropathol Appl Neurobiol. 46(6):588-601 (2020)
- Eede P, Obst J, Benke E, Yvon-Durocher G, Richard BC, Gimber N, Schmoranzer J, Böddrich A, Wanker EE, Prokop S, Heppner FL. Interleukin-12/23 deficiency differentially affects pathology in male and female Alzheimer's disease-like mice. EMBO Rep. 21(3):e48530 (2020)
- Lehmann M, Lukonin I, Noé F, Schmoranzer J, Clementi C, Loerke D, Haucke V. Nanoscale coupling of endocytic pit growth and stability. Sci Adv. 5(11):eaax5775 (2019)
- Brosig A, Fuchs J, Ipek F, Kroon C, Schrötter S, Vadhvani M, Polyzou A, Ledderose J, van Diepen M, Holzhütter HG, Trimbuch T, Gimber N, Schmoranzer J, Lieberam I, Rosenmund C, Spahn C, Scheerer P, Szczepek M, Leondaritis G, Eickholt BJ. The Axonal Membrane Protein PRG2 Inhibits PTEN and Directs Growth to Branches. Cell Rep. 29(7):2028-2040.e8 (2019)
- Brockmann MM, Maglione M, Willmes CG, Stumpf A, Bouazza BA, Velasquez LM, Grauel MK, Beed P, Lehmann M, Gimber N, Schmoranzer J, Sigrist SJ, Rosenmund C, Schmitz D. RIM-BP2 primes synaptic vesicles via recruitment of Munc13-1 at hippocampal mossy fiber synapses. Elife. 8:e43243 (2019)
- Giesecke T, Himmerkus N, Leipziger J, Bleich M, Koshimizu TA, Fähling M, Smorodchenko A, Shpak J, Knappe C, Isermann J, Ayasse N, Kawahara K, Schmoranzer J, Gimber N, Paliege A, Bachmann S, Mutig K. Vasopressin Increases Urinary Acidification via V1a Receptors in Collecting Duct Intercalated Cells. J Am Soc Nephrol. 30(6):946-961 (2019)
- Song K, Gras C, Capin G, Gimber N, Lehmann M, Mohd S, Puchkov D, Rödiger M, Wilhelmi I, Daumke O, Schmoranzer J, Schürmann A, Krauss M. A SEPT1-based scaffold is required for Golgi integrity and function. J Cell Sci. 132(3):jcs225557 (2019)
- Houtman J, Freitag K, Gimber N, Schmoranzer J, Heppner FL, Jendrach M. Beclin1-driven autophagy modulates the inflammatory response of microglia via NLRP3. EMBO J. 38(4):e99430 (2019)
- Kath C, Goni-Oliver P, Müller R, Schultz C, Haucke V, Eickholt B, Schmoranzer J. PTEN suppresses axon outgrowth by down-regulating the level of detyrosinated microtubules. PLoS One. 13(4):e0193257 (2018)
- Gao, M., Maraspini, R., Beutel, O., Zehtabian, A., Eickholt, B., Honigmann, A. and Ewers, H. Expansion stimulated emission depletion Microscopy (ExSTED). ACS Nano 12, 4178-4185 (2018)
- Schöneberg, J., M. Lehmann, A. Ullrich, Y. Posor, W.-T. Lo, G. Lichtner, J. Schmoranzer, V. Haucke, and F. Noé. Lipid-Mediated PX-BAR Domain Recruitment Couples Local Membrane Constriction to Endocytic Vesicle Fission. Nature Comm. 8: 15873 (2017)
- Albrecht, D., Winterflood, C. M., Sadeghi, M., Tschager, T., Noé, F., and Ewers, H. Nanoscopic compartmentalization of membrane protein motion at the axon initial segment. J Cell Biol. 215, 37–46 (2016)
- Lehmann, M., G. Lichtner, H. Klenz, and J. Schmoranzer. Novel Organic Dyes for Multicolor Localization-Based Super-Resolution Microscopy. J. of Biophotonics 9 (1–2): 161–70 (2016)
- Gimber, N., G. Tadeus, T. Maritzen, J. Schmoranzer, and V. Haucke. Diffusional Spread and Confinement of Newly Exocytosed Synaptic Vesicle Proteins. Nature Comm. 6: 8392 (2015)
- Lehmann, M., B. Gottschalk, D. Puchkov, P. Schmieder, S. Schwagerus, C.P.R. Hackenberger, V. Haucke, and J. Schmoranzer. Multicolor Caged DSTORM Resolves the Ultrastructure of Synaptic Vesicles in the Brain. Angew. Ch. Int. Ed. 54 (45): 13230–35 (2015)
- Mikhaylova M., Cloin, B.M.C., Finan, K., van den Berg, R., Teeuw, J., Kijanka, M.M., Sokolowski, M., Katrukha, E.A., Maidorn, M. Opazo, F., Moutel, S., Vantard, M. Perez, F.,van Bergen en Henegouwen, P.M.P., Hoogenraad, C.C., Ewers, H., and Kapitein, L.C. Resolving bundled microtubules using anti-tubulin nanobodies. Nat Comm 11; 6:7933 (2015)
- Kaplan, C., Jing, B., Winterflood, C.W., Bridges, A.A., Occhipinti, P., Schmied, J., Grinhagens, S., Gronemeyer, T., Tinnefeld, P., Gladfelter, A.S., Ries, J. and Ewers, H. The absolute arrangement of subunits in cytoskeletal septin filaments in cells measured by fluorescence microscopy. Nano Letters 15(6):3859-64 (2015)
- Winterflood, C.M., Platonova, E., Albrecht, D. and Ewers H. Simple and robust dual color 3D superresolution microscopy by combined spectral-demixing and bi-plane imaging. Biophysical Journal 109 (1) pp:3-6 (2015)
- Koo SJ, Kochlamazashvili G, Rost B, Puchkov D, Gimber N, Lehmann M, Tadeus G, Schmoranzer J, Rosenmund C, Haucke V, Maritzen T. Vesicular Synaptobrevin/VAMP2 Levels Guarded by AP180 Control Efficient Neurotransmission. Neuron. 88(2):330-44. doi: 10.1016/j.neuron.2015.08.034 (2015)
- Sakaba T, Kononenko NL, Bacetic J, Pechstein A, Schmoranzer J, Yao L, Barth H, Shupliakov O, Kobler O, Aktories K, Haucke V. Fast neurotransmitter release regulated by the endocytic scaffold intersectin. Proc Natl Acad Sci U S A. 110(20):8266-71 (2013)
- Podufall J., Tian R., Knoche E., Jung N., Lampe A., Wichmann C., Zhang Y.Q., Schmoranzer J., Sigrist S.J., Haucke V., “A presynaptic role for the cytomatrix protein GIT in synaptic vesicle recycling”, *manuscript in preparation* 16. Hucho T., Suckow V., Joseph E.K., Kuhn J., Schmoranzer J., Dina O.A., Chen X., Karst M., Bernateck M., Levine J.D., Ropers H.-H. Ca++/CaMKII switches nociceptor sensitizing stimuli into desensitizing stimuli. J. Neurochemistry 123(4):589-601 (2012)
- Lampe A., Haucke V., Sigrist S., Heilemann M., Schmoranzer J. Multi-color direct STORM with red-emitting carbocyanines. Biology of the Cell 104(4):229-37 (2012)
- Schmoranzer J, Fawcett JP, Segura M, Vallee RB, Pawson T, Gundersen GG. Par3 and dynein associate to regulate local microtubule dynamics and centrosome orientation during migration. Current Biology 19(13):1065-74 (2009)
- Bartolini F, Moseley JB, Schmoranzer J, Cassimeris L, Goode BL, Gundersen GG. The formin mDia2 stabilizes microtubules independently of its actin nucleation activity. J. Cell Biology 181(3):523-36 (2008)
- Gundersen GG, Wen Y, Eng CH, Schmoranzer J, Canbera-Poch N, Morris EJS, Chen M, Gomes E. Regulation of microtubules by Rho GTPases in migrating cells, Novartis Found Symp 269:106-16 (2005)
- Wen Y, Eng CH, Schmoranzer J, Canbera-Poch N, Morris EJS, Chen M, Wallar BJ, Alberts AS, Gundersen GG. EB1 and APC interact with mDia to selectively stabilize microtubules downstream of Rho and promote cell migration. Nature Cell Biology 6(9):820-30 (2004)
- Schmoranzer J, Kreitzer G and Simon SM. Migrating fibroblasts perform polarized and micro-tubule-dependent exocytosis towards the leading edge. J. Cell Science 116(22):4513-19 (2003)
- Schmoranzer J and Simon SM. Role of microtubules in transport and fusion of post-Golgi vesicles at the plasma membrane. Mol. Biol. Cell 4(4):1558-69 (2003)
- Kreitzer G, Schmoranzer J, Gan Y, Low SH, Zhang Z, Weimbs T, Simon SM and Rodriguez-Boulan E. Three dimensional analysis of post-Golgi carrier exocytosis in polarized epithelial cells. Nature Cell Biology 5(2):126-36 (2003)
- Lampson MA, Schmoranzer J, Zeigerer A, Simon SM, and McGraw TE. Insulin-regulated release from the endosomal recycling compartment is regulated by budding of specialized vesicles. Mol. Biol. Cell, 12(11):3489-501 (2001)